
Vaughn Jackson, PhD
Emeritus Professor
Locations
- Biochemistry
BSB 351
Contact Information
Education
Biography
Dr. Jackson received his Doctorate degree in Biochemistry at the University of Iowa in 1972. His postdoctoral research was at the University of Iowa from 1972 to 1976 and at the MRC Lab of Molecular Biology, Cambridge, U.K. from 1976 to 1978.
He was a research associate scientist at the University of Iowa from 1978 to 1982 and joined the faculty at the Medical College of Wisconsin in 1982.
Research Interests
Transcription within a eucaryotic cell is not a process in which RNA polymerase can freely access the DNA for either initiation or elongation of the transcript. The DNA is associated with highly basic proteins called histones which function to condense the DNA in an organized manner and to facilitate accessibility of the DNA under regulated conditions. Four of these histones, H3, H2A, H2B, H4 coil the DNA into a left-handed supercoil and form a particle called a nucleosome. The DNA is organized into a tandem array of these nucleosomes. Because of the strong binding energies between these proteins and DNA, specific cellular mechanisms are required to access DNA for transcription or replication. We are involved in a characterization of the following potential mechanisms:
1. The role of metabolic modifications such as acetylation. A common characteristic of active genes is the presence of histones that are highly acetylated.
2. The role of topological stress in DNA. A characteristic of the transcriptional process is the formation of positive stress in advance of the RNA polymerase and negative stress in its wake.
3. The role of accessory proteins in modulating histone-DNA interactions. Several recently characterized proteins have been shown to function in either an ATP dependent or independent process to disrupt these interactions.
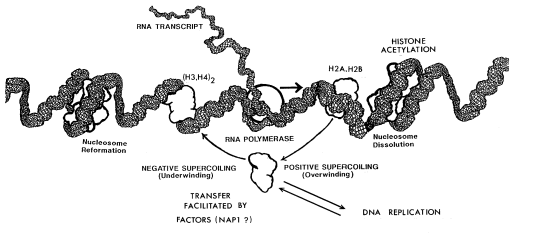
The figure illustrates a model of transcription which encompasses these three potential mechanisms. Laboratory procedures for testing this model involve both in vivo and in vitro experimentation. The in vivo procedures require exposure of tissue culture cells to radioactive and or density-labeled precursors in order to label proteins and or DNA under conditions which alter either replication or transcription. The dynamics of histone-DNA interactions can be studied by this approach. The in vitro procedures involve the assembly of individual components into a well defined transcription system that is designed to minimize variables. By a manipulation of the in vitro conditions, it is possible to simulate the in vivo observations. In this way, we are able to study specific regulatory mechanism in detail.
Publications
-
(White RH, Keberlein M, Jackson V.) Biochemistry. 2012 Oct 16;51(41):8173-88 PMID: 23003102 09/26/2012
-
The nucleosome family: dynamic and growing.
(Zlatanova J, Bishop TC, Victor JM, Jackson V, van Holde K.) Structure. 2009 Feb 13;17(2):160-71 PMID: 19217387 02/17/2009
-
(Peterson S, Jackson V.) Biochemistry. 2008 Jul 08;47(27):7053-65 PMID: 18543948 06/12/2008
-
(Peterson S, Danowit R, Wunsch A, Jackson V.) Biochemistry. 2007 Jul 24;46(29):8634-46 PMID: 17595058 06/28/2007
-
(Sun HY, McNally MT, Jackson VE, Grossberg SE.) J Virol Methods. 2006 Nov;137(2):304-8 PMID: 16920200 PMCID: PMC2575689 SCOPUS ID: 2-s2.0-33748714112 08/22/2006
-
The FK506-binding protein, Fpr4, is an acidic histone chaperone.
(Xiao H, Jackson V, Lei M.) FEBS Lett. 2006 Aug 07;580(18):4357-64 PMID: 16846601 07/19/2006
-
(Wunsch A, Jackson V.) Biochemistry. 2005 Dec 13;44(49):16351-64 PMID: 16331996 12/08/2005
-
(Levchenko V, Jackson B, Jackson V.) Biochemistry. 2005 Apr 12;44(14):5357-72 PMID: 15807529 04/06/2005
-
(Levchenko V, Jackson V.) Biochemistry. 2004 Mar 09;43(9):2359-72 PMID: 14992573 03/03/2004
-
In vitro studies on the maintenance of transcription-induced stress by histones and polyamines.
(Peng HF, Jackson V.) J Biol Chem. 2000 Jan 07;275(1):657-68 PMID: 10617664 01/05/2000
-
Formaldehyde cross-linking for studying nucleosomal dynamics.
(Jackson V.) Methods. 1999 Feb;17(2):125-39 PMID: 10075891 03/17/1999
-
(Peng HF, Jackson V.) Biochemistry. 1997 Oct 07;36(40):12371-82 PMID: 9315878 10/07/1997

